Posts Tagged ‘what is a species’
Microbe biogeography: the distribution, dispersal and evolution of the littlest organisms
EDIT: This post was selected as a runner up for the PLoS ONE Blog Pick of the Month. Thanks, PLoS! Check out the winner: Greg Laden’s great post on how the victims of Vesuvius died.
![]() In any high school biology class1, we learn that isolation is key to the evolution of species. For example, take Australia, where an array of marsupials such as koalas and kangaroos reproduce like no other animals on the planet. Isolation on a continental island allowed ancestral marsupials to evolve gestation via pouch, a trait which was retained as these animals later evolved into multiple (cuddly) species. In other words: an event that happened in the past resulted in the organisms we see today, or the history of a species influences its current form and life history.
In any high school biology class1, we learn that isolation is key to the evolution of species. For example, take Australia, where an array of marsupials such as koalas and kangaroos reproduce like no other animals on the planet. Isolation on a continental island allowed ancestral marsupials to evolve gestation via pouch, a trait which was retained as these animals later evolved into multiple (cuddly) species. In other words: an event that happened in the past resulted in the organisms we see today, or the history of a species influences its current form and life history.
We attribute the distribution of species on this planet, also known as biogeography, to these sorts of historical events. Organisms evolved, and continue to evolve, the way they do due to historical circumstances out of their control, creating the biodiversity of our world. The idea of biogeography is generally attributed to Lamarck, and throughout the late-18th and early-19th centuries (pre-Darwin, mind you), scientists suggested many reasons for the non-uniform distribution of organisms, with Lyell summing up these historical factors as a combination of environment and dispersal through migration, passive (e.g. seeds carried in the wind) or active (e.g. elephants walking across the plain).

It's hard to find images for these kinds of posts, ok? Cut me some slack. By the way, image (c) Hannah Waters 2010
However, not all organisms seemed to fit this pattern. Scientists at this time observed that, while polar bears were limited to the arctic and monkeys to warm climes, organisms such as fungi, sponges, algae and lichens were far more ubiquitous. The botanist Kurt Sprengel, in summary of a common thought, wrote that organisms of “lower organization” must have greater ability to disperse, allowing them to colonize more broadly and thrive where “circumstances propitious to their production occur.” (For a full history, see Maureen O’Malley’s commentary in Nature Reviews Microbiology.)
In 1934, the Dutch biologist Lourens Baas-Becking revived this idea, with the thought that the typical explanations of biogeography do not fit with the world of microorganisms. He saw the same species of microbe living in different places on the globe and in variable environments. Thus, he posited that historical factors such as isolation and environment could not be the forces determining microbial distribution, but rather that “everything is everywhere; the environment selects.” The small size and abundance of many microbe species allowed them to be easily dispersed in water, on wind, on the bodies of animals, spreading them all over the planet. Many microbes can also lie dormant for a long time until conditions improve, or until the “environment selects” them. This would, in effect, create what’s been termed a “seed bank” of microbes, where all microbes are in all environments at the same time, lying in wait for environmental conditions to favor their proliferation.
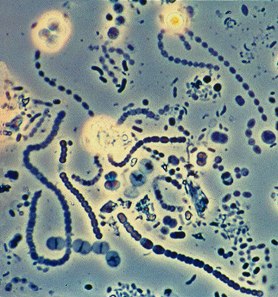
A generic microbial community. Source: Frank Dazzo, Center for Microbial Ecology, Michigan State University
For most of the 20th century, this so-called “Bass-Becking Hypothesis” was widely accepted, but in the past few decades has been hotly debated. In 2004, Tom Fenchel and Bland Finlay compiled a literature review in Bioscience in favor of the hypothesis, arguing that “habitat properties alone are needed to explain the presence of a given microbe, and historical factors are irrelevant.” They reviewed studies which showed the ubiquity of microbe species with fewer habitat requirements (generalists, if you will), as well as microbe species that are environmentally specific but are found in their preferred habitats on many continents. Of note is a 1997 Oikos study that they themselves published, wherein they found 20 living microbe species in a lake sample. Upon altering conditions (such as food source, temperature, acidity, and oxygen levels), they were able to revive an additional 110 species – evidence supporting the idea of a “seed bank” of microbes. The authors do note that this theory may only apply to the most common microbe species, since not all are able to dessicate and revive – but perhaps this ability is what made them so widespread in the first place.
One caveat with this study is that the authors advocate for a phenotypic analysis of microbes. While the ability to study DNA was a huge benefit to the field of microbiology, the authors do not agree that this is useful due to the wide genetic variability even within a single microbial population, and thus rely on morphology to describe species instead of genetic analysis. A 2006 review, including genetic analyses, found that things aren’t so cut and dry. The authors cite a number of studies showing reproducible genetic differences within microbe species even along a 10-meter transect in a marsh. In two hot springs thousands of kilometers apart, despite living in the same environment, two species of bacteria (Synechococcus and Sulfolobus) showed significant genetic differences. This shows that isolation alone can affect genes, and thus ultimately species, “overwhelming any effect of environmental factors.”
Both reviews note that there is not enough data out there to draw strong conclusions; the 2006 study was relying on 10 articles alone to determine distance and environmental signficance. To me the differences in these studies come down to how one defines a “species.” Typically, we differentiate species based on an organism’s ability to produce fertile offspring with another – if they can, they are the same species. (There are many caveats to the “species problem” beyond my scope right now. For a really thorough write-up, see this post from the Wild Muse.) However, most microbes reproduce via cell division, and genes can be transferred horizontally despite “species” boundaries. So how do we even define a microbial species in the first place? If we’re looking at evolution alone, it would seem that genetic differences even within microbes that are commonly described as the same species morphologically would be meaningful, as these genetic differences put them on the path to become novel species.
One major question that the idea of “everything is everywhere” brings up is: how do microbes evolve in the first place? If these organisms are relatively free from the external pressures of isolation and environment, going locally extinct or reviving based on their surrounding conditions, evolution must take an incredibly long time.
I could not find a paper on biogeography and microbial evolution; however, a paper in PLoS published in April 2010 looked at the biogeography and large-scale evolution of phytoplankton in the ocean. In light of questions I’m asking here, oceanic plankton and microbe communities are very similar. They are both small organisms primarily dispersed passively, by ocean currents in the case of plankton. The ocean hosts a wide variety of environments, and plankton are also generally considered to be everywhere at once. While it is not ideal, I will use this planktonic model to look at biogeography and evolution in a more specific system. (Well, as specific as you can get with the ocean…)
Just as the determinants of microbial biogeography haven’t been concluded, the same is true of plankton. In this study, the authors sampled planktonic communities in two very different ocean environments: subtropical/tropical oceans, characterized by similar conditions throughout a wide geographical range, highly stratified ocean layers, and nutrient-poor surface waters, and sub-Arctic waters, characterized by high vertical mixing and high nutrient levels across the water column. They compared 250-ml samples pairwise from each of the oceanic habitats and found that the planktonic communities were “strikingly dissimilar.” However, when they increased their sample size 100-fold to 25 liters, they found that these contrasting ocean environments shared 76% of their total species pool! This effect is surely found in many microbial studies: when comparing diversity between smaller plots, you are more likely to find a difference. But an increase in plot size, even within the same environment, will find more similarities. (Which is a more meaningful measurement is another question… I’d be happy to hear your comments on that one.)
To look at the evolution of phytoplankton, the authors took core samples from four distinct geographic environments and then identified fossil diatom species within from 240 million years ago to the present, generating “community assemblages” of diatoms through time. They then compared these communities assemblages with environmental factors: global CO2 concentrations and oceanic upwelling strength. The authors found that, despite “local determinants such as regional current systems, terrestrial nutrient inputs, atmospheric deposition, physical mixing, etc.,” global climate measures largely predicted the diatom community assemblage, with many species recovering after local extinction. That’s right: even after the extinction of a species, when preferable environmental conditions returned, so did the diatom.
This study provides a clue regarding the importance of environmental conditions to the global distribution of abundant, passively dispersed organisms. What is also interesting is that the same diatom species were found again and again over the course of 240 million years. Their ability for high dispersal and recovery of species enables planktonic communities to evolve “slowly and gradually” over time.
But clearly they have evolved: plankton (and microbes) are incredibly diverse clades. The question to look at now is how is evolution driven in highly dispersed organisms?
And thus, as usual, they are the tiniest organisms that force us to broaden our view on basic tenets of biology. Just as horizontal gene transfer did for traditional natural selection, now microbial dispersal does for the evolution of species.
It does give me a great deal of hope regarding life on this planet: the possibility that there is a cache of microbes waiting around for the perfect conditions, even ones not suitable for us. As my father, Dennis P. Waters (who needs a blog), once put it, “As long as there’s bacteria, there’s hope.”
1That is, in one where evolution is taught at all…
Cermeño, P., de Vargas, C., Abrantes, F., & Falkowski, P. (2010). Phytoplankton Biogeography and Community Stability in the Ocean PLoS ONE, 5 (4) DOI: 10.1371/journal.pone.0010037
Fenchel, T., & Finlay, B. (2004). The Ubiquity of Small Species: Patterns of Local and Global Diversity BioScience, 54 (8) DOI: 10.1641/0006-3568(2004)054[0777:TUOSSP]2.0.CO;2
Martiny, J., Bohannan, B., Brown, J., Colwell, R., Fuhrman, J., Green, J., Horner-Devine, M., Kane, M., Krumins, J., Kuske, C., Morin, P., Naeem, S., Øvreås, L., Reysenbach, A., Smith, V., & Staley, J. (2006). Microbial biogeography: putting microorganisms on the map Nature Reviews Microbiology, 4 (2), 102-112 DOI: 10.1038/nrmicro1341
O’Malley, M. (2007). The nineteenth century roots of ‘everything is everywhere’ Nature Reviews Microbiology, 5 (8), 647-651 DOI: 10.1038/nrmicro1711
Quote of the Day: Lamarck on Species
From Lamarck’s Zoological Philosophy (1809)
The farther we advance in our knowledge of the various organised bodies which cover almost every part of the earth’s surface, the greater becomes our difficulty in determining what should be regarded as a species, and still more in finding the boundaries and distinctions of genera.
According as the productions of nature are collected and our museums grow richer, we see nearly all gaps filled up and the lines of demarcation effaced. We find ourselves reduced to an arbitrary decision which sometimes leads us to take the smallest differences of varieties and erect them into what we call species, and sometimes leads us to describe as a variety of some species slightly differing individuals which others regard as constituting a separate species.
Just as true now as then.
And for your information: Lamarck’s full name is Jean Baptiste Pierre Antoine de Monet Lamarck.
Invasive species corrupt DNA, not just ecosystems (Fitzpatrick et al., PNAS 2010)
I rarely think about how invasive species affect genetics. It’s always in terms of ecosystems or species: invasive brown tree snakes gobbling up birds and lizards in Guam, or zebra mussels overwhelming and altering the environment of the Great Lakes. How one species outcompetes and replaces another, changing the natural system. This is partly because many of the common examples are of predator-prey relationships, where the two species are very distantly related and could never breed, thus keeping genetics out of the picture. But what about situations where the introduced animal and native animal are similar?
This gets us into the muddy waters of what defines a species. For sexually reproducing organisms, a species is the group of animals with whom one can exchange genetic material via reproduction, or, in other words, can produce fertile offspring. To distinguish one species from another under this definition, a scientist would need a pretty wide worldview. How else could he know that a squirrel from England could not mate with a US squirrel if it tried? And the honest answer is: he can’t. (Unless he collected squirrels from around the world and tried to mate them all with one another… but that’s a lot of work.) Thus, species are often also defined based on location or geography, despite the fact that maybe they could mate if they had access to one another. But what are the chances that a squirrel will swim across the Atlantic for a new girlfriend?
 And there’s where invasive species fit in. In a paper published this week in PNAS out of Knoxville, TN, Lexington, KY, and UC Davis, scientists studied the Salinas Valley in central California, where salamanders from Texas and New Mexico had been introduced in the 1950s for use as bait by fisherman. These salamanders, the Barred Tiger Salamanders (Ambystoma tigrinum mavortium) had been defined as a separate species from the threatened native California Tiger Salamanders (Ambystoma californiense), as their populations had been living apart for 3-10 million years, and thus it was unlikely that they were still genetically similar enough to mate. But – alas – this assumption was wrong. The invasive salamanders have been mating with the native species for the last 50 years, producing hybrids which are able to mate with either species and one another. The question: is this hybridization significantly changing the DNA of the native species?
And there’s where invasive species fit in. In a paper published this week in PNAS out of Knoxville, TN, Lexington, KY, and UC Davis, scientists studied the Salinas Valley in central California, where salamanders from Texas and New Mexico had been introduced in the 1950s for use as bait by fisherman. These salamanders, the Barred Tiger Salamanders (Ambystoma tigrinum mavortium) had been defined as a separate species from the threatened native California Tiger Salamanders (Ambystoma californiense), as their populations had been living apart for 3-10 million years, and thus it was unlikely that they were still genetically similar enough to mate. But – alas – this assumption was wrong. The invasive salamanders have been mating with the native species for the last 50 years, producing hybrids which are able to mate with either species and one another. The question: is this hybridization significantly changing the DNA of the native species?
To investigate this question, the researchers identified an introduction site at a pond in central California, and took samples of over 200 salamanders (by clipping the end of their tail and immediately releasing them) at this site and others within a region 200 km north. Using salamanders of each species from non-invaded ranges, they determined the baseline genetic makeup of each species.
They scanned the genomes of the sampled salamanders (say it 10 times fast) for 68 genetic markers to see if any of the invasive genes had “taken over” the native genes. They saw no real difference in 65 of these species — that is, the salamanders retained their native genes. However, they saw a drastic increase in 3 of the genes. In the figure below, taken from their paper (click on image for larger size), the little “thermometers” measure the DNA differences at different sites, native in white and invasive in black, with the introduction site indicated by the red arrow. The upper left (A) shows the big picture: of the 68 gene markers studied total, invasive genes are only apparent at the introduction site. The other 3 boxes (B, C, D) show the three genes that have spread — and as you can see, they have spread far and deep, despite their invisibility overall (A). The authors were thorough: they tested whether this pattern was due to either sampling error or random genetic drift without natural selection, and neither of these biases accounted for the pattern of these 3 genes.
The function of these genes is unknown. However, by studying the behavior of the animals, it seems they are related to reproduction. The hybrids have larger larvae with greater survival and develop more quickly, ever hastening their dispersal. This raises a few questions:
1. If these invasive genes are helping survival, then who cares if they invaded? It is easy to look at this as actually beneficial to the threatened native salamanders. However, it has unknown impacts on the surrounding ecosystem. These bigger larvae eat a lot more, impacting the populations of their prey species through indirect effects of the invasion. A change in the abundance of one species affects all others – what seems to be an immediate benefit can be incredibly harmful in the long run.
2. How do we define a species? The native salamanders are a threatened species. If they have received genes that increase their numbers through hybridization, is this a comeback? Are they still A. californiense? Do these 3 genes alone make them A. mavortium? Are they an entirely new species? Is it possible to stop the invasion of these genes throughout the state without killing off a threatened species?
I don’t have the answers to these questions. We human beings are drawn to classification: we want to put all of the animals in neat little piles and call it fin. But the truth is that species are eternally evolving — as Peter and Rosemary Grant have shown with their Galapagos finches, most recently in November 2009. The monkeys that live on one side of a jungle can have a different genetic makeup than the ones on the other side even if they can still mate.
Clearly the introduction of these salamanders, which was just an innocent attempt to raise some bait locally, has had unforeseen impacts on the ecosystem. Humans’ ability to travel has meant that we are bringing animals together that have not evolved to live together, or have evolved apart millions of years ago. In some ways it feels like what is done is done — and I am not enough of an expert on habitat restoration to tell you otherwise. But try little things: wash the mud off of your boots before you go hiking in another state or country, don’t release your foreign pets locally (as my roommate Erinrose and I have been tempted to do with our pet turtle, Nicolas Cage), volunteer at your local wildlife refuge. Biodiversity is important.
How can we save our planet?
Fitzpatrick, B., Johnson, J., Kump, D., Smith, J., Voss, S., & Shaffer, H. (2010). Rapid spread of invasive genes into a threatened native species Proceedings of the National Academy of Sciences, 107 (8), 3606-3610 DOI: 10.1073/pnas.0911802107







