Posts Tagged ‘life histories’
Microbe biogeography: the distribution, dispersal and evolution of the littlest organisms
EDIT: This post was selected as a runner up for the PLoS ONE Blog Pick of the Month. Thanks, PLoS! Check out the winner: Greg Laden’s great post on how the victims of Vesuvius died.
![]() In any high school biology class1, we learn that isolation is key to the evolution of species. For example, take Australia, where an array of marsupials such as koalas and kangaroos reproduce like no other animals on the planet. Isolation on a continental island allowed ancestral marsupials to evolve gestation via pouch, a trait which was retained as these animals later evolved into multiple (cuddly) species. In other words: an event that happened in the past resulted in the organisms we see today, or the history of a species influences its current form and life history.
In any high school biology class1, we learn that isolation is key to the evolution of species. For example, take Australia, where an array of marsupials such as koalas and kangaroos reproduce like no other animals on the planet. Isolation on a continental island allowed ancestral marsupials to evolve gestation via pouch, a trait which was retained as these animals later evolved into multiple (cuddly) species. In other words: an event that happened in the past resulted in the organisms we see today, or the history of a species influences its current form and life history.
We attribute the distribution of species on this planet, also known as biogeography, to these sorts of historical events. Organisms evolved, and continue to evolve, the way they do due to historical circumstances out of their control, creating the biodiversity of our world. The idea of biogeography is generally attributed to Lamarck, and throughout the late-18th and early-19th centuries (pre-Darwin, mind you), scientists suggested many reasons for the non-uniform distribution of organisms, with Lyell summing up these historical factors as a combination of environment and dispersal through migration, passive (e.g. seeds carried in the wind) or active (e.g. elephants walking across the plain).

It's hard to find images for these kinds of posts, ok? Cut me some slack. By the way, image (c) Hannah Waters 2010
However, not all organisms seemed to fit this pattern. Scientists at this time observed that, while polar bears were limited to the arctic and monkeys to warm climes, organisms such as fungi, sponges, algae and lichens were far more ubiquitous. The botanist Kurt Sprengel, in summary of a common thought, wrote that organisms of “lower organization” must have greater ability to disperse, allowing them to colonize more broadly and thrive where “circumstances propitious to their production occur.” (For a full history, see Maureen O’Malley’s commentary in Nature Reviews Microbiology.)
In 1934, the Dutch biologist Lourens Baas-Becking revived this idea, with the thought that the typical explanations of biogeography do not fit with the world of microorganisms. He saw the same species of microbe living in different places on the globe and in variable environments. Thus, he posited that historical factors such as isolation and environment could not be the forces determining microbial distribution, but rather that “everything is everywhere; the environment selects.” The small size and abundance of many microbe species allowed them to be easily dispersed in water, on wind, on the bodies of animals, spreading them all over the planet. Many microbes can also lie dormant for a long time until conditions improve, or until the “environment selects” them. This would, in effect, create what’s been termed a “seed bank” of microbes, where all microbes are in all environments at the same time, lying in wait for environmental conditions to favor their proliferation.
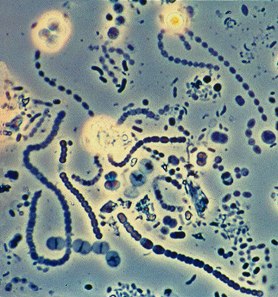
A generic microbial community. Source: Frank Dazzo, Center for Microbial Ecology, Michigan State University
For most of the 20th century, this so-called “Bass-Becking Hypothesis” was widely accepted, but in the past few decades has been hotly debated. In 2004, Tom Fenchel and Bland Finlay compiled a literature review in Bioscience in favor of the hypothesis, arguing that “habitat properties alone are needed to explain the presence of a given microbe, and historical factors are irrelevant.” They reviewed studies which showed the ubiquity of microbe species with fewer habitat requirements (generalists, if you will), as well as microbe species that are environmentally specific but are found in their preferred habitats on many continents. Of note is a 1997 Oikos study that they themselves published, wherein they found 20 living microbe species in a lake sample. Upon altering conditions (such as food source, temperature, acidity, and oxygen levels), they were able to revive an additional 110 species – evidence supporting the idea of a “seed bank” of microbes. The authors do note that this theory may only apply to the most common microbe species, since not all are able to dessicate and revive – but perhaps this ability is what made them so widespread in the first place.
One caveat with this study is that the authors advocate for a phenotypic analysis of microbes. While the ability to study DNA was a huge benefit to the field of microbiology, the authors do not agree that this is useful due to the wide genetic variability even within a single microbial population, and thus rely on morphology to describe species instead of genetic analysis. A 2006 review, including genetic analyses, found that things aren’t so cut and dry. The authors cite a number of studies showing reproducible genetic differences within microbe species even along a 10-meter transect in a marsh. In two hot springs thousands of kilometers apart, despite living in the same environment, two species of bacteria (Synechococcus and Sulfolobus) showed significant genetic differences. This shows that isolation alone can affect genes, and thus ultimately species, “overwhelming any effect of environmental factors.”
Both reviews note that there is not enough data out there to draw strong conclusions; the 2006 study was relying on 10 articles alone to determine distance and environmental signficance. To me the differences in these studies come down to how one defines a “species.” Typically, we differentiate species based on an organism’s ability to produce fertile offspring with another – if they can, they are the same species. (There are many caveats to the “species problem” beyond my scope right now. For a really thorough write-up, see this post from the Wild Muse.) However, most microbes reproduce via cell division, and genes can be transferred horizontally despite “species” boundaries. So how do we even define a microbial species in the first place? If we’re looking at evolution alone, it would seem that genetic differences even within microbes that are commonly described as the same species morphologically would be meaningful, as these genetic differences put them on the path to become novel species.
One major question that the idea of “everything is everywhere” brings up is: how do microbes evolve in the first place? If these organisms are relatively free from the external pressures of isolation and environment, going locally extinct or reviving based on their surrounding conditions, evolution must take an incredibly long time.
I could not find a paper on biogeography and microbial evolution; however, a paper in PLoS published in April 2010 looked at the biogeography and large-scale evolution of phytoplankton in the ocean. In light of questions I’m asking here, oceanic plankton and microbe communities are very similar. They are both small organisms primarily dispersed passively, by ocean currents in the case of plankton. The ocean hosts a wide variety of environments, and plankton are also generally considered to be everywhere at once. While it is not ideal, I will use this planktonic model to look at biogeography and evolution in a more specific system. (Well, as specific as you can get with the ocean…)
Just as the determinants of microbial biogeography haven’t been concluded, the same is true of plankton. In this study, the authors sampled planktonic communities in two very different ocean environments: subtropical/tropical oceans, characterized by similar conditions throughout a wide geographical range, highly stratified ocean layers, and nutrient-poor surface waters, and sub-Arctic waters, characterized by high vertical mixing and high nutrient levels across the water column. They compared 250-ml samples pairwise from each of the oceanic habitats and found that the planktonic communities were “strikingly dissimilar.” However, when they increased their sample size 100-fold to 25 liters, they found that these contrasting ocean environments shared 76% of their total species pool! This effect is surely found in many microbial studies: when comparing diversity between smaller plots, you are more likely to find a difference. But an increase in plot size, even within the same environment, will find more similarities. (Which is a more meaningful measurement is another question… I’d be happy to hear your comments on that one.)
To look at the evolution of phytoplankton, the authors took core samples from four distinct geographic environments and then identified fossil diatom species within from 240 million years ago to the present, generating “community assemblages” of diatoms through time. They then compared these communities assemblages with environmental factors: global CO2 concentrations and oceanic upwelling strength. The authors found that, despite “local determinants such as regional current systems, terrestrial nutrient inputs, atmospheric deposition, physical mixing, etc.,” global climate measures largely predicted the diatom community assemblage, with many species recovering after local extinction. That’s right: even after the extinction of a species, when preferable environmental conditions returned, so did the diatom.
This study provides a clue regarding the importance of environmental conditions to the global distribution of abundant, passively dispersed organisms. What is also interesting is that the same diatom species were found again and again over the course of 240 million years. Their ability for high dispersal and recovery of species enables planktonic communities to evolve “slowly and gradually” over time.
But clearly they have evolved: plankton (and microbes) are incredibly diverse clades. The question to look at now is how is evolution driven in highly dispersed organisms?
And thus, as usual, they are the tiniest organisms that force us to broaden our view on basic tenets of biology. Just as horizontal gene transfer did for traditional natural selection, now microbial dispersal does for the evolution of species.
It does give me a great deal of hope regarding life on this planet: the possibility that there is a cache of microbes waiting around for the perfect conditions, even ones not suitable for us. As my father, Dennis P. Waters (who needs a blog), once put it, “As long as there’s bacteria, there’s hope.”
1That is, in one where evolution is taught at all…
Cermeño, P., de Vargas, C., Abrantes, F., & Falkowski, P. (2010). Phytoplankton Biogeography and Community Stability in the Ocean PLoS ONE, 5 (4) DOI: 10.1371/journal.pone.0010037
Fenchel, T., & Finlay, B. (2004). The Ubiquity of Small Species: Patterns of Local and Global Diversity BioScience, 54 (8) DOI: 10.1641/0006-3568(2004)054[0777:TUOSSP]2.0.CO;2
Martiny, J., Bohannan, B., Brown, J., Colwell, R., Fuhrman, J., Green, J., Horner-Devine, M., Kane, M., Krumins, J., Kuske, C., Morin, P., Naeem, S., Øvreås, L., Reysenbach, A., Smith, V., & Staley, J. (2006). Microbial biogeography: putting microorganisms on the map Nature Reviews Microbiology, 4 (2), 102-112 DOI: 10.1038/nrmicro1341
O’Malley, M. (2007). The nineteenth century roots of ‘everything is everywhere’ Nature Reviews Microbiology, 5 (8), 647-651 DOI: 10.1038/nrmicro1711
Photosynthetic Evolution: how 2 organisms gained or lost the ability to eat sunshine
9/13/2010 Update
New research has come out that changes the story told below. Wägele et al. sequenced cDNA transcripts from RNA produced by slugs dependent on their plastids alone, and did not find the RNA to produce the proteins for plastid use in any meaningful quantity. But the slugs aren’t just using the proteins they got upon original ingestion; we simply do not know what is happening. For a thorough write-up, I highly recommend this post from The Spandrel Shop entitled “Solar Sea Slugs: part plant, part animal… or not?”
Original post (01/20/2010):

The variety of protist life -- from Finlay, B.J. & Esteban, G.F. Protist, 149, 155-165 (via the British Society of Protist Biology).
Biologists and taxonomists love to put organisms into categories to help us organize the complicated living world. I grew up on the 5 kingdom system of classification: plants, animals, fungi, bacteria and protists. The first four categories seemed simple enough, but the term “protists” always confused me. This kingdom seemed to be a dumping ground for all the single-celled organisms that we didn’t know what to do with, ones that had evolved so far from their ancestors that their origins were unknown.
I’ve stumbled upon two fascinating articles about such animals that are out of place. The first is about microorganisms that were once photosynthetic — and thus evolved with the cyanobacteria and plants — but no longer go through photosynthesis. The second is about a sea slug that has developed the ability to photosynthesize, or harvest energy from the sun. Imagine stumbling upon these animals for the first time. The former would be placed in the animal kingdom due to its heterotrophy, or tendency to get its nutrients from food. The latter would be baffling: is it some freak highly-organized plant, or an anomalous, energy-producing animal? Thanks to our increasingly understanding of evolution, scientists have been able to figure out where these strange creatures came from.
The first story is about evolution driven by competition for food between many species of microorganisms. In terms of energy acquisition, I usually think about two categories of organisms: the heterotrophs, which get their energy from eating, and the autotrophs, which create their own energy from outside inputs such as the sun. But there are also the mixotrophs, which are able to do both. To be a mixotroph! How wonderful would it be to get energy from standing under the sun if you wanted, but you could also be able to eat a hearty meal for the same gain! On first thought, this appears to be the best life strategy, allowing an organism to get energy whatever way is easiest at the time.
Scientists from the University of Potsdam in Germany and the Austrian Academy of Scientists propose that, in some circumstances, having both strategies may be too much (open access paper here). In order to be a mixotroph, the organism needs to fit all the machinery required for both processes inside its single cell, increasing its size. A bigger mixotrophic cell not only loses access to smaller food items, such as ultramicobacteria, but also has to compete with larger heterotrophic organisms, which are solely dedicated to eating and thus can do so more efficiently. These scientists created a model showing how, under low-light conditions, it would be energenically beneficial for mixotrophs to be rid of their chloroplasts and other organelles needed for photosynthesis so that they could become physically smaller and have access to the smaller foods out of the range of other heterotrophs. Thus, we have animal-like organisms that evolved from plants.
 The second story is about a sea slug, Elysia chlorotica, which has gained the ability to photosynthesize. It did not evolve this trait in the traditional sense, but rather picked it up from another organism. The slug’s green color is not self-made, but is present due to its collection of chloroplasts, the photosynthetic center of a cell, from its prey. Due to an unknown mechanism, the slug is able to hoard only the chloroplasts of its algal food source Vaucheria litoria. Not only that, but it uses these chloroplasts to go through photosynthesis itself, which it can continue to do 5 months after it last ate V. litoria. (And this is a slug that only lives for 10 months total.)
The second story is about a sea slug, Elysia chlorotica, which has gained the ability to photosynthesize. It did not evolve this trait in the traditional sense, but rather picked it up from another organism. The slug’s green color is not self-made, but is present due to its collection of chloroplasts, the photosynthetic center of a cell, from its prey. Due to an unknown mechanism, the slug is able to hoard only the chloroplasts of its algal food source Vaucheria litoria. Not only that, but it uses these chloroplasts to go through photosynthesis itself, which it can continue to do 5 months after it last ate V. litoria. (And this is a slug that only lives for 10 months total.)
This is not as simple as it sounds, however. You need more than chloroplasts to photosynthesize; you also need genes to encode all the specialized proteins needed to make sunlight into energy! The big question regarding these slugs was: where did they get these genes? Scientists working together from Maine, Korea, Iowa and Texas (paper here) compared sections of the nuclear DNA between the slug and its algal food and found identical segments, suggesting that the slug had not evolved these genes on their own, but had acquired them through horizontal gene transfer, or a transfer of DNA from an origin other than one’s own parent. In this case, they suggest, a segment of the algae’s DNA broke off and joined the slug’s own DNA, an incredibly rare event. This gene acquisition was so beneficial that it spread through the population, causing E. chlorotica all over the oceans to hoard the chloroplasts of their prey. And there ya have it, folks: a photosynthetic animal.
Ain’t this a wonderful world we live in?
Learning about the histories of these organisms makes me drool, thinking about the uncategorized protists out there. What kind of stories do they have to tell?
de Castro, F., Gaedke, U., & Boenigk, J. (2009). Reverse Evolution: Driving Forces Behind the Loss of Acquired Photosynthetic Traits PLoS ONE, 4 (12) DOI: 10.1371/journal.pone.0008465
Rumpho, M., Worful, J., Lee, J., Kannan, K., Tyler, M., Bhattacharya, D., Moustafa, A., & Manhart, J. (2008). Horizontal gene transfer of the algal nuclear gene psbO to the photosynthetic sea slug Elysia chlorotica Proceedings of the National Academy of Sciences, 105 (46), 17867-17871 DOI: 10.1073/pnas.0804968105






